Selective Vulnerability to Neurodegenerative Diseases
We use mouse genetic tools to understand how specific brain regions are affected by neurodegenerative processes. We also are interested in understanding how poor sleep affects the onset and progression of neurodegenerative disease. We address these problems by developing models of diseases and tools to study them. One of our main technical focuses in recent years is to study how gene regulation in specific cell types is affected by neurodegenerative disease and sleep loss, using genetically encoded tools and next generation sequencing techniques. Manipulation of gene sequence and activity with CRISPR/Cas based methods is also a strength of the lab.
Selective vulnerability to neurodegeneration
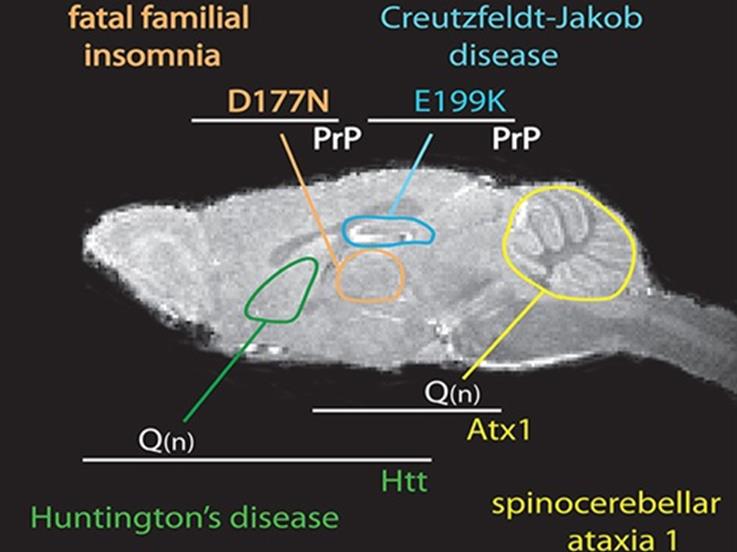
Neurodegenerative diseases cause stereotypical signs based on the specific brain region that is targeted. Our lab addresses the question of why neurodegenerative diseases target specific brain regions, at least initially, a feature known as selective vulnerability. For example, brain regions important for motor control are severely damaged in Huntington’s disease, while brain regions important for memory are severely damaged in Alzheimer’s disease. We try to understand how unaffected regions resist disease with the hope that these secrets can be transferred to vulnerable brain regions.
Genetic neurodegenerative diseases are especially enigmatic because all cells carry the disease inducing mutation and yet most resist the disease. Moreover, these mutations are expressed for decades before disease sets in. How does the brain resist the toxic protein for so long and why does it eventually fail?
Questions
We have two basic questions regarding selective vulnerability:
1) Does a given brain cell respond differently to different protein misfolding stressors and
2) Do different cell types respond to the same stressor differently?
We hypothesize that the effectiveness of the protein quality control machinery differs between different cell types due to different compilations of machinery components and clients and these differences strongly influence selective vulnerability.
To study the phenotypes of specific cell types in the brain prior to disease onset, we have employed a technique that enables the isolation of mRNAs from the cells of interest. This is accomplished by expressing in specific cell types a modified ribosome protein (called RiboTag) that carries a molecular handle which we use to capture translating ribosomes and determine which proteins the cells are attempting to make.
Our work with RiboTag revealed that cells that are thought to resist disease show dramatic changes early, while cells that are severely affected late in disease show no response early. These surprising results inspired us to create new technique that enables us to study multiple layers of gene regulation, from chromatin modulation to translation, all from specific cell types of complex tissues.